Key words: miRNAs, IsomiRs
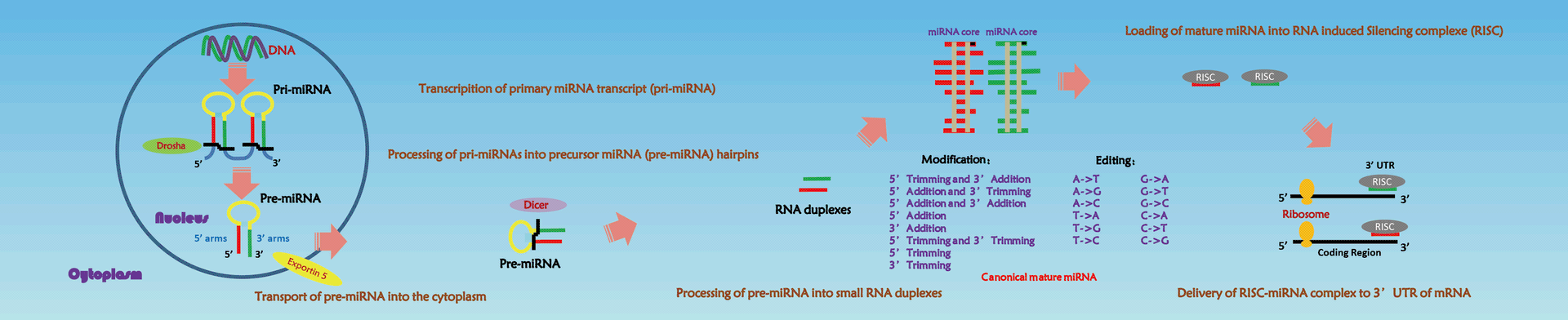
Mature microRNAs (miRNAs), average length of 22 nucleotides, are expressed ubiquitously and regulate many essential biological processes and pathological conditions mainly via post transcriptional silencing through mRNA decay and/or translational repression [1]. Recently, Next-Generation Sequencing (NGS) technology has revealed that miRNA frequently exhibit differences from their corresponding reference mature sequences, generating multiple variants specifically abbreviated as microRNA isoforms (isomiRs) [2]. Isoforms are generated either due to imprecise miRNAs precursor cleavage, terminal trimming or the addition of non-templated nucleotide via exoribonuclease or nucleotidyl transferase activity. IsomiRs are functional in influence the miRNA stability, sub-cellular localization and target selection [3-5].
IsomiRs are detected by using our previously published algorithm CPSS [6]. IsomiR Bank was implemented in PHP + MySQL + JavaScript + R and can be accessed without any registration. It is the first integrative resource that contains the sequence and expression of isomiRs. In total, 2,727 samples (Small RNA NGS data downloaded for ArrayExpress) from eight species (Arabidopsis thaliana, Drosophila melanogaster, Danio rerio, Homo sapiens, Mus musculus, Oryza sativa, Solanum lycopersicum and Zea mays) were analyzed by using CPSS. We have stored 308,919 isomiRs from 4,706 mature miRNAs in our IsomiR Bank database. Furthermore, we have provided the figures in order to illustrate the effects of isomiRs on targets selection and GO analysis for these affected targets.