GAS (Greedy AUC Stepwise) is a classification framework, which is successfully applied to binary classifier. In 0/1 classification problem, GAS maximizes AUC of ROC curve with pregiven number of variables automatically. The goal is to solve a K-Sparse problem. For a K-sparse problem, we assume the number of non-zero features should be less than K. GAS is different from regularity-penalized methods such as compressive sensing and LASSO. When a potential model is given, such as SVM (with any given kernel function) and logistic regression, GAS will find the best K features using a special way of greedy searching, then give the final model with the maximize AUC. The strategy of GAS is similar to forward selection, which only adds one variable that is not already in the model and increases the value of AUC. If GAS fails to find out the solution with K variables, it will output the model that generates the maximum AUC instead For each step with maximum allowed numbers. The algorithm is described as follows.
Greed AUC Stepwise
Require initialize selected variable stack \( V \), and AUC stack \( A \). Ensure Generating proper logistic model
Initial STEP
\( i\leftarrow 0 \), compute NULL model: \( AUC\leftarrow auc_0 \), A.push(\( auc_0 \))
STEP 1\( i++ \)
\( auc_i=max^{*}_j \{auc_{ij}\},\quad\forall j\in I/V \); where \( auc_{ij} \) is the AUC of LRM with \( V\cup j \) variables
A.push(\( auc_i \)); V.push(\( argmax^{*}_{j\in I/V}\{auc_{ij}\} \));STEP 2
\( auc_i < auc_{i-1}; i-- \)
A.pull(), V.pull()
Goto Step 1 but change the definition of \( max^{*} \) as the second (or third, fourth, etc ) largest number.
if \( max^{*} \) can not be well-definedbreak
EndIf
Else
Goto Step 1
EndIf
OUTPUT final model
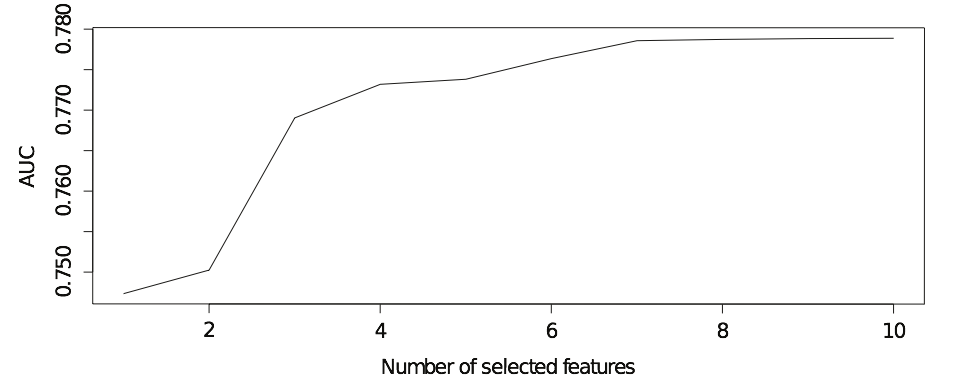
A typical solution path of GAS is a strictly increasing step function:

One may notice GAS will add the variables into the model in every step.
The performance in [1] can be seen from the following ROC curve.
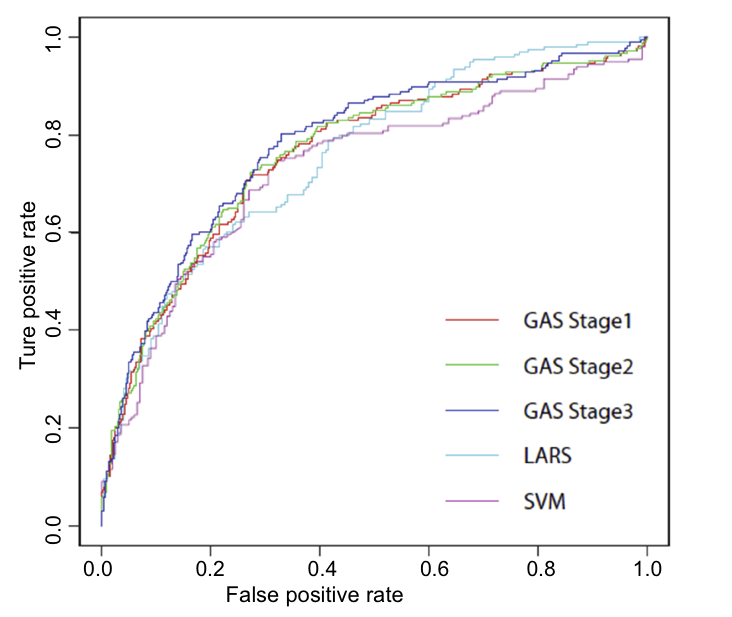
In this case, GAS promote the result of tranditional GLM (family = logistic).
| Models | Sn | Sp | MCC | AC | AUC |
|---|---|---|---|---|---|
| GAS (STAGE1) | 0.360 | 0.914 | 0.329 | 0.636 | 0.751 |
| GAS (STAGE2) | 0.445 | 0.909 | 0.399 | 0.676 | 0.752 |
| GAS (STAGE3) | 0.530 | 0.859 | 0.411 | 0.693 | 0.758 |
| LARS (GLMNET) | 0.570 | 0.808 | 0.389 | 0.688 | 0.741 |
| SVM (Linear) | 0.620 | 0.823 | 0.355 | 0.673 | 0.718 |
Ref
[1] Besides, one may notice that AUC is equivalent to Gini coefficient, i.e. \[ Gini=2\times{}AUC-1 \] So, the loss function used in GAS could be modified to Gini coefficient or Pearson-\( \chi^2 \) statistics. If likelihood function with penalty is used, GAS is actually a modification of forward selection.
[2] Zhang, Yuanwei, Liangwen Zhong, Bo Xu, Yifan Yang, Rongjun Ban, Jun Zhu, Howard J. Cooke, QiaoMei Hao, and Qinghua Shi. “SpermatogenesisOnline 1.0: a resource for spermatogenesis based on manual literature curation and genome-wide data mining.” Nucleic acids research 41, no. D1 (2013): D1055-D1062.