General information
| SG000221 | |||||||||||||||||
| Gja1 | |||||||||||||||||
| AU042049, AW546267, Cnx43, Cx43, Cx43alpha1, Gja-1, Npm1, connexin43 | |||||||||||||||||
| Mus musculus | |||||||||||||||||
| 10090 | |||||||||||||||||
| ENSMUSG00000050953 | |||||||||||||||||
| ENSMUSP00000064536 ENSMUSP00000151596 ENSMUSP00000151603 ENSMUSP00000151647 ENSMUSP00000151974 ENSMUSP00000151620 | |||||||||||||||||
Gap junction protein, alpha 1 [Mus musculus (house mouse)] | |||||||||||||||||
| |||||||||||||||||
Reviewed functional gene |
Functional information
| premeiotic meiotic postmeiotic | |
| Leydig cell Sertoli cell Spermatogonia | |
|
1. Abstract 2. Abstract 3. Abstract 4. Abstract 5. Abstract 6. Abstract | |
| |
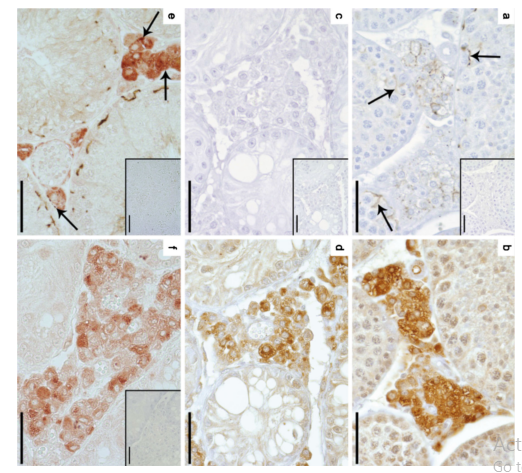
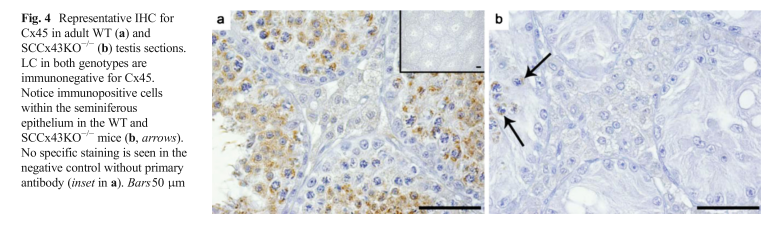
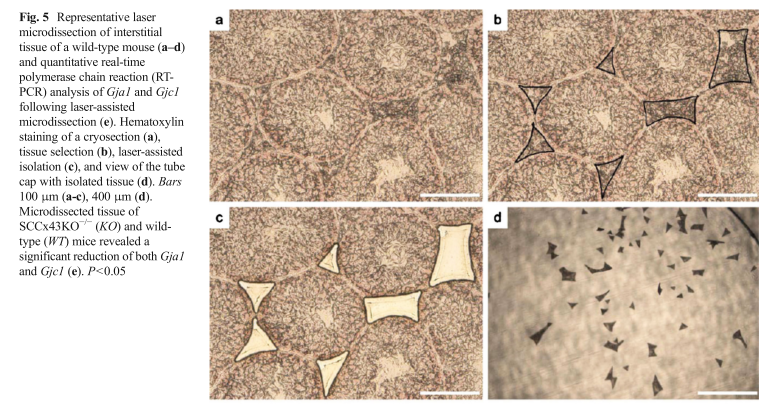
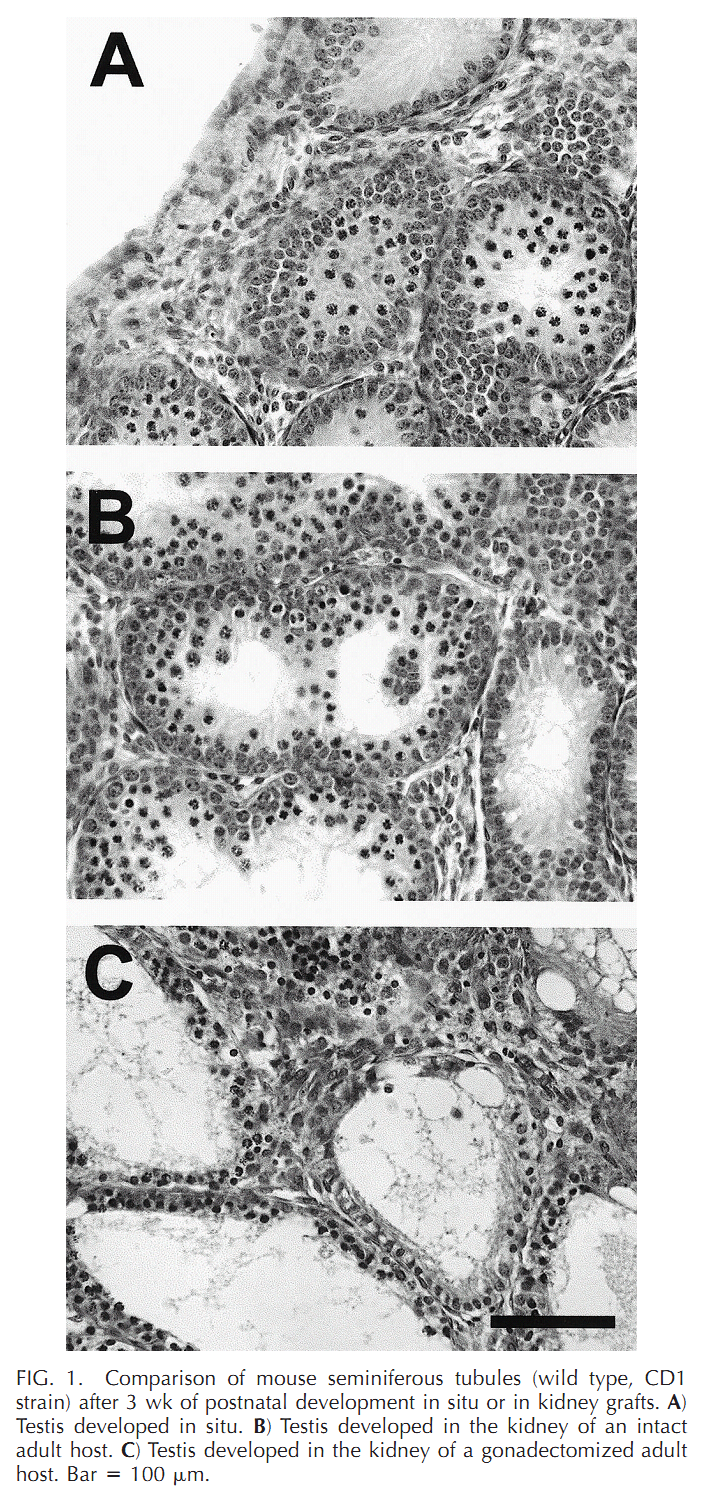
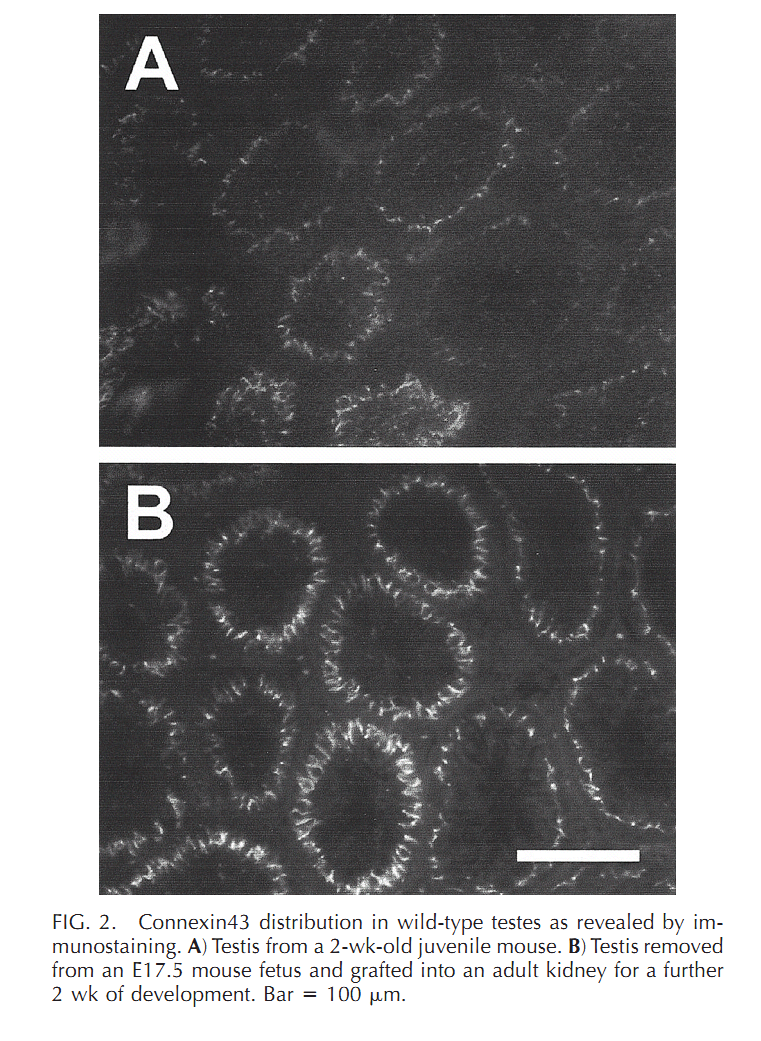
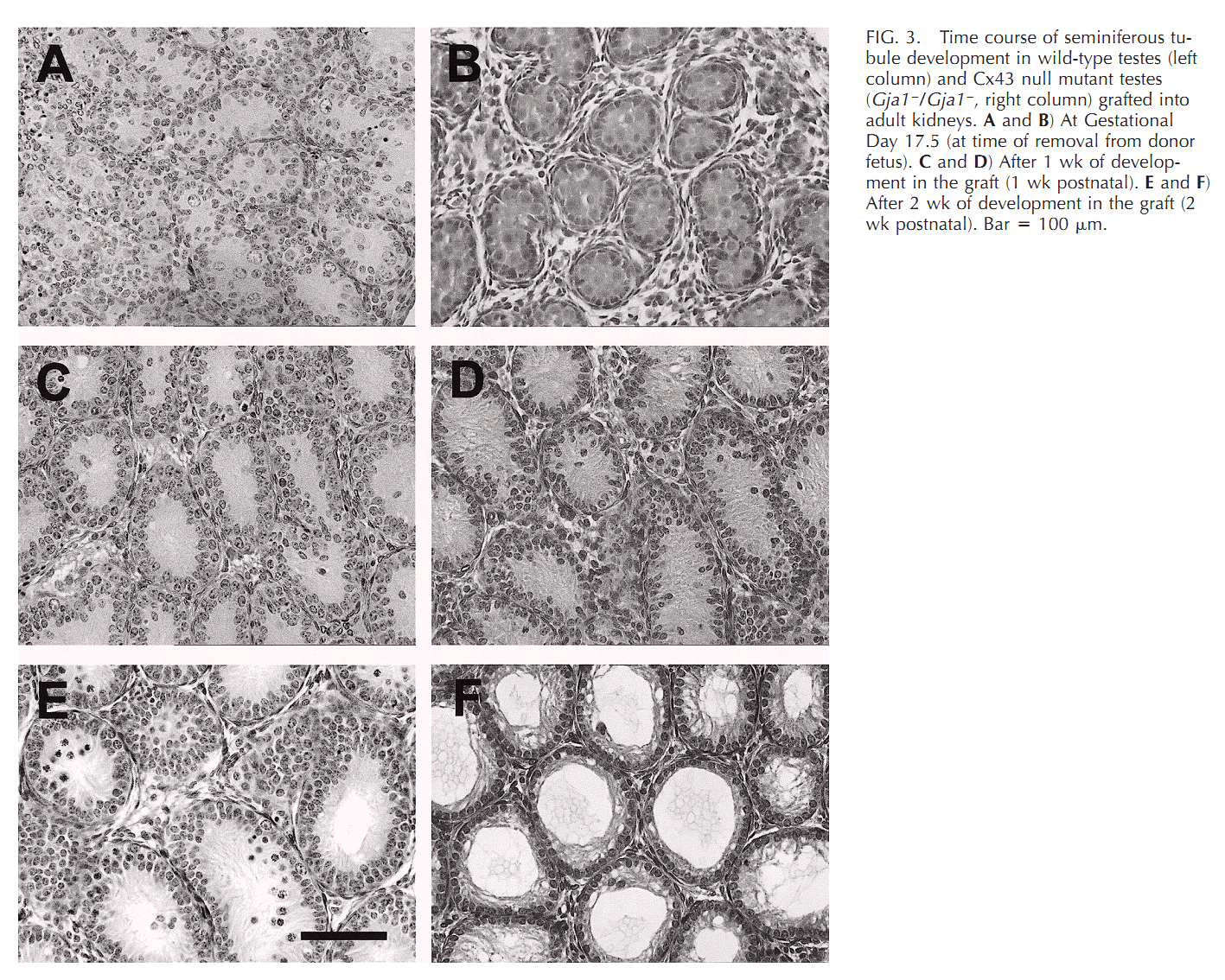
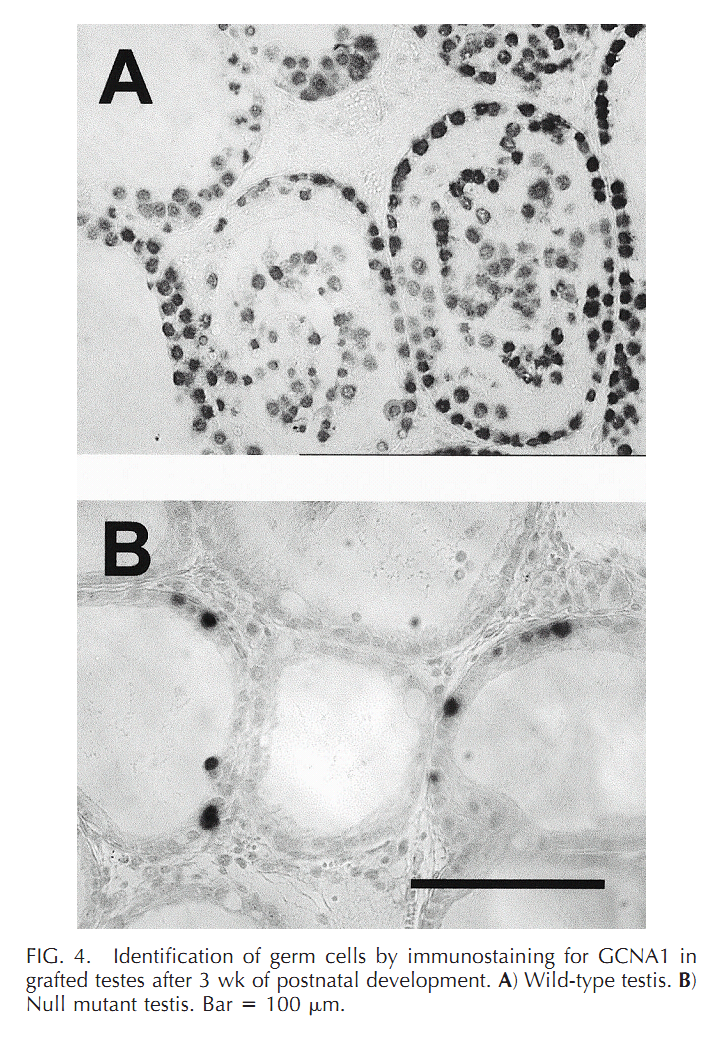
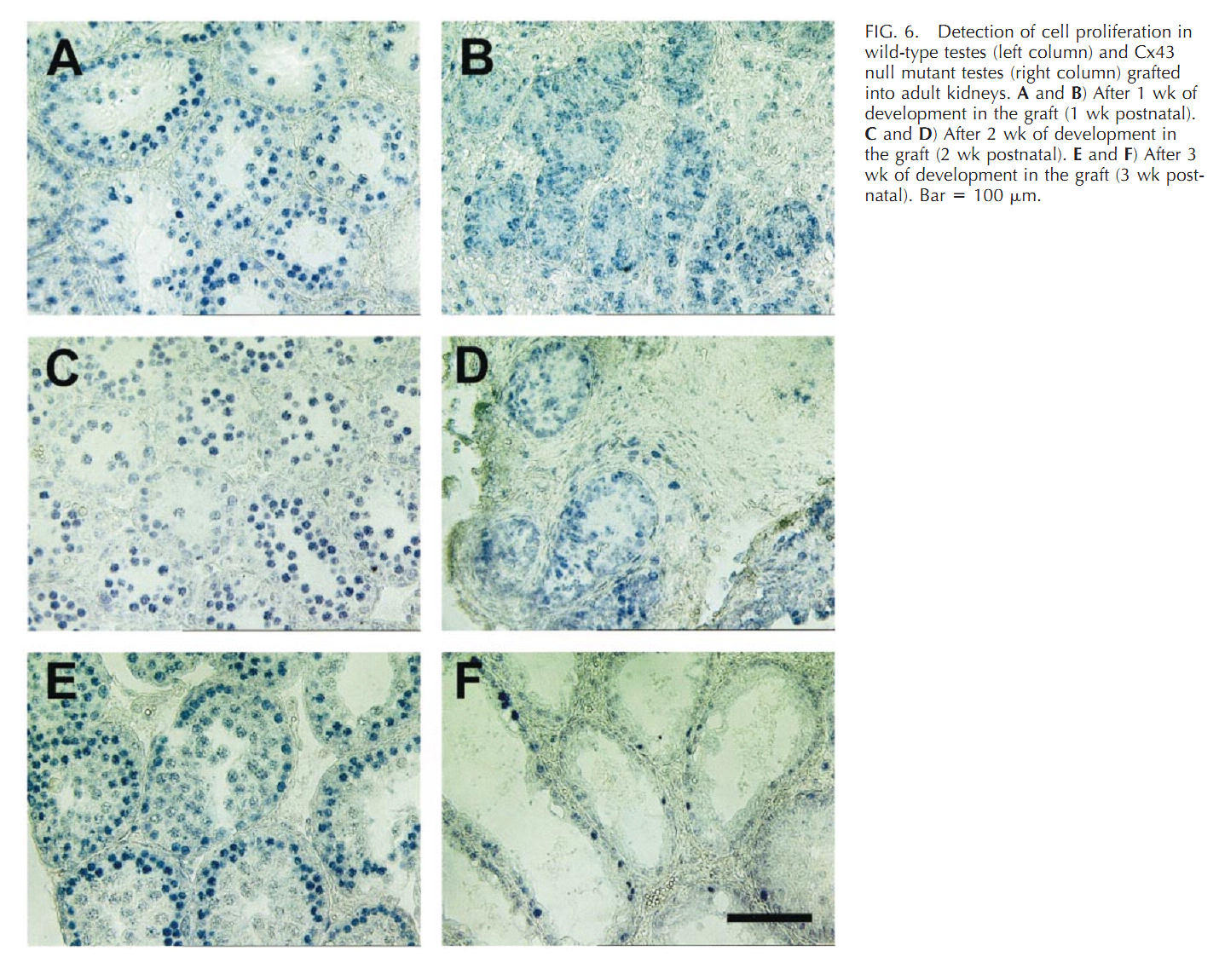
| Cx43 (Gja1) is the predominant testicular Cx and is expressed by the Sertoli cells, germ cells , peritubular cells, and. Cx43 has been demonstrated to be involved in SC proliferation and differentiation; it plays an essential role in the initiation and maintenance of spermatogenesis and is possibly involved in the steroidogenesis of interstitial Leydig cells. | |
|
ReactomeID: R-MMU-196025 Formation of annular gap junctions | |
| SCO |
Expression and location
| Expressed highest in skin | |
| Expressed highest in spermatogonium | |
| View detail | |
| View detail | |
| Cell membrane |
Mutation in human orthology
|
1000G (Phase 3): 381 ESP6500 (SI-V2): 38 ExAC (r0.3.1): 217 dbSNP(Build 147): 1018 |
|||||||
|
Chinese health control (254): 1 European health control (283): 2 Chinese patients (168): |
|||||||
|
Other information
|
Click the right sign for more information
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| P23242 |